Inference with xgboost
Source:vignettes/articles/Inference-with-xgboost.Rmd
Inference-with-xgboost.RmdThis article contains a workflow in R to analyze a data set using xgboost to get insights that can help a consultant make important business decisions.
knitr::opts_chunk$set(warning = FALSE, message = FALSE)
library(pacman)
library(tidyverse); library(EIX); library(validata); p_load(TidyConsultant)
set.seed(1)Inspect data set
We will use the HR_data from the EIX package. Let’s inspect the variables using the validata package.
HR_data
#> satisfaction_level last_evaluation number_project average_montly_hours
#> <num> <num> <int> <int>
#> 1: 0.38 0.53 2 157
#> 2: 0.80 0.86 5 262
#> 3: 0.11 0.88 7 272
#> 4: 0.72 0.87 5 223
#> 5: 0.37 0.52 2 159
#> ---
#> 14995: 0.40 0.57 2 151
#> 14996: 0.37 0.48 2 160
#> 14997: 0.37 0.53 2 143
#> 14998: 0.11 0.96 6 280
#> 14999: 0.37 0.52 2 158
#> time_spend_company Work_accident left promotion_last_5years sales
#> <int> <int> <int> <int> <fctr>
#> 1: 3 0 1 0 sales
#> 2: 6 0 1 0 sales
#> 3: 4 0 1 0 sales
#> 4: 5 0 1 0 sales
#> 5: 3 0 1 0 sales
#> ---
#> 14995: 3 0 1 0 support
#> 14996: 3 0 1 0 support
#> 14997: 3 0 1 0 support
#> 14998: 4 0 1 0 support
#> 14999: 3 0 1 0 support
#> salary
#> <fctr>
#> 1: low
#> 2: medium
#> 3: medium
#> 4: low
#> 5: low
#> ---
#> 14995: low
#> 14996: low
#> 14997: low
#> 14998: low
#> 14999: low
HR_data %>%
diagnose_category(max_distinct = 100)
#> # A tibble: 13 × 4
#> column level n ratio
#> <chr> <fct> <int> <dbl>
#> 1 sales sales 4140 0.276
#> 2 sales technical 2720 0.181
#> 3 sales support 2229 0.149
#> 4 sales IT 1227 0.0818
#> 5 sales product_mng 902 0.0601
#> 6 sales marketing 858 0.0572
#> 7 sales RandD 787 0.0525
#> 8 sales accounting 767 0.0511
#> 9 sales hr 739 0.0493
#> 10 sales management 630 0.0420
#> 11 salary low 7316 0.488
#> 12 salary medium 6446 0.430
#> 13 salary high 1237 0.0825
HR_data %>%
diagnose_numeric()
#> # A tibble: 8 × 10
#> variables zeros minus infs min mean max `|x|<=1 (ratio)` integer_ratio
#> <chr> <chr> <chr> <chr> <int> <int> <int> <chr> <chr>
#> 1 satisfacti… 0 (0… 0 (0… 0 (0… 0 0 1 14999 (100%) 111 (1%)
#> 2 last_evalu… 0 (0… 0 (0… 0 (0… 0 0 1 14999 (100%) 283 (2%)
#> 3 number_pro… 0 (0… 0 (0… 0 (0… 2 3 7 0 (0%) 14999 (100%)
#> 4 average_mo… 0 (0… 0 (0… 0 (0… 96 201 310 0 (0%) 14999 (100%)
#> 5 time_spend… 0 (0… 0 (0… 0 (0… 2 3 10 0 (0%) 14999 (100%)
#> 6 Work_accid… 1283… 0 (0… 0 (0… 0 0 1 14999 (100%) 14999 (100%)
#> 7 left 1142… 0 (0… 0 (0… 0 0 1 14999 (100%) 14999 (100%)
#> 8 promotion_… 1468… 0 (0… 0 (0… 0 0 1 14999 (100%) 14999 (100%)
#> # ℹ 1 more variable: mode <chr>xgboost binary classification model
Create dummy variables out of the Sales and Salary column. We will
predict whether an employee left the company using xgboost. For this
reason, set left = 1 to the first level of the factor, so
it will be treated as the event class. A high predicted indicates a
label of the event class.
HR_data %>%
framecleaner::create_dummies() %>%
framecleaner::set_fct(left, first_level = "1") -> hr1Create the model using xgboost. Since the goal of this model is to run inference using trees, we want to set tree_depth to 2 to make easily-interpretable trees.
When the model is run, feature importance on the full data is printed. Also the data is split into train and test, where the accuracy is calculated on a test set. Since this is a binary classification problem, a confusion matrix is output along with binary metrics.
hr1 %>%
tidy_formula(left) -> hrf
hr1 %>%
tidy_xgboost(hrf, tree_depth = 2L, trees = 100L, mtry = .75) -> xg1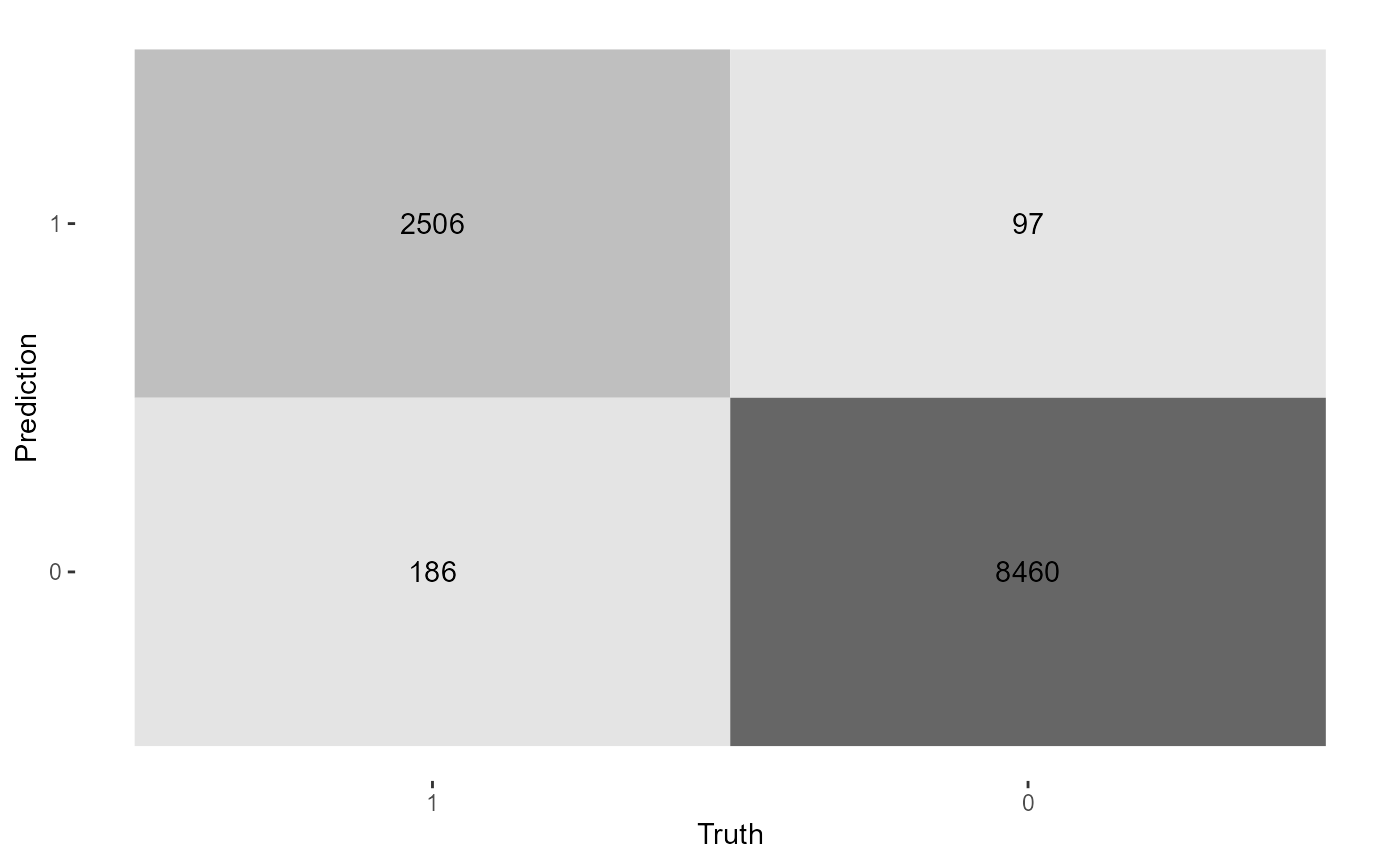
#> # A tibble: 15 × 3
#> .metric .estimate .formula
#> <chr> <dbl> <chr>
#> 1 accuracy 0.958 TP + TN / total
#> 2 kap 0.883 NA
#> 3 sens 0.871 TP / actually P
#> 4 spec 0.986 TN / actually N
#> 5 ppv 0.953 TP / predicted P
#> 6 npv 0.960 TN / predicted N
#> 7 mcc 0.884 NA
#> 8 j_index 0.857 NA
#> 9 bal_accuracy 0.928 sens + spec / 2
#> 10 detection_prevalence 0.221 predicted P / total
#> 11 precision 0.953 PPV, 1-FDR
#> 12 recall 0.871 sens, TPR
#> 13 f_meas 0.910 HM(ppv, sens)
#> 14 baseline_accuracy 0.758 majority class / total
#> 15 roc_auc 0.977 NA
This line will save the tree structure of the model as a table.
xg1 %>%
xgboost::xgb.model.dt.tree(model = .) -> xg_trees
xg_trees
#> Tree Node ID Feature Split Yes No Missing
#> <int> <int> <char> <char> <num> <char> <char> <char>
#> 1: 0 0 0-0 average_montly_hours 288.00 0-1 0-2 0-2
#> 2: 0 1 0-1 satisfaction_level 0.47 0-3 0-4 0-4
#> 3: 0 2 0-2 Leaf NA <NA> <NA> <NA>
#> 4: 0 3 0-3 Leaf NA <NA> <NA> <NA>
#> 5: 0 4 0-4 Leaf NA <NA> <NA> <NA>
#> ---
#> 608: 99 2 99-2 average_montly_hours 162.00 99-5 99-6 99-6
#> 609: 99 3 99-3 Leaf NA <NA> <NA> <NA>
#> 610: 99 4 99-4 Leaf NA <NA> <NA> <NA>
#> 611: 99 5 99-5 Leaf NA <NA> <NA> <NA>
#> 612: 99 6 99-6 Leaf NA <NA> <NA> <NA>
#> Gain Cover
#> <num> <num>
#> 1: 8.012269e+02 2056.34204
#> 2: 2.794244e+03 2010.99219
#> 3: 2.076119e-01 45.34982
#> 4: 9.449380e-02 532.40680
#> 5: -3.902787e-02 1478.58533
#> ---
#> 608: 6.358565e+01 1055.31921
#> 609: -8.300844e-02 14.49944
#> 610: -5.473283e-02 18.97126
#> 611: 2.040233e-02 342.93545
#> 612: -5.765188e-03 712.38373Let’s plot the first tree and interpret the table output. For tree=0, the root feature (node=0) is satisfaction level, which is split at value .465. Is satisfaction_level < .465? If Yes, observations go left to node 1, if no, observations go right to node 2. Na values would go to node 1 if present. The quality of the split is represented by its Gain: 3123, the improvement in training loss.
xgboost::xgb.plot.tree(model = xg1, trees = 0)The quality in the leaves is the prediction for observations in those leaves represented by log odds. To interpret them as probabilities, use the function below. Importantly, a log odds of 0 is a 0.5 probability.
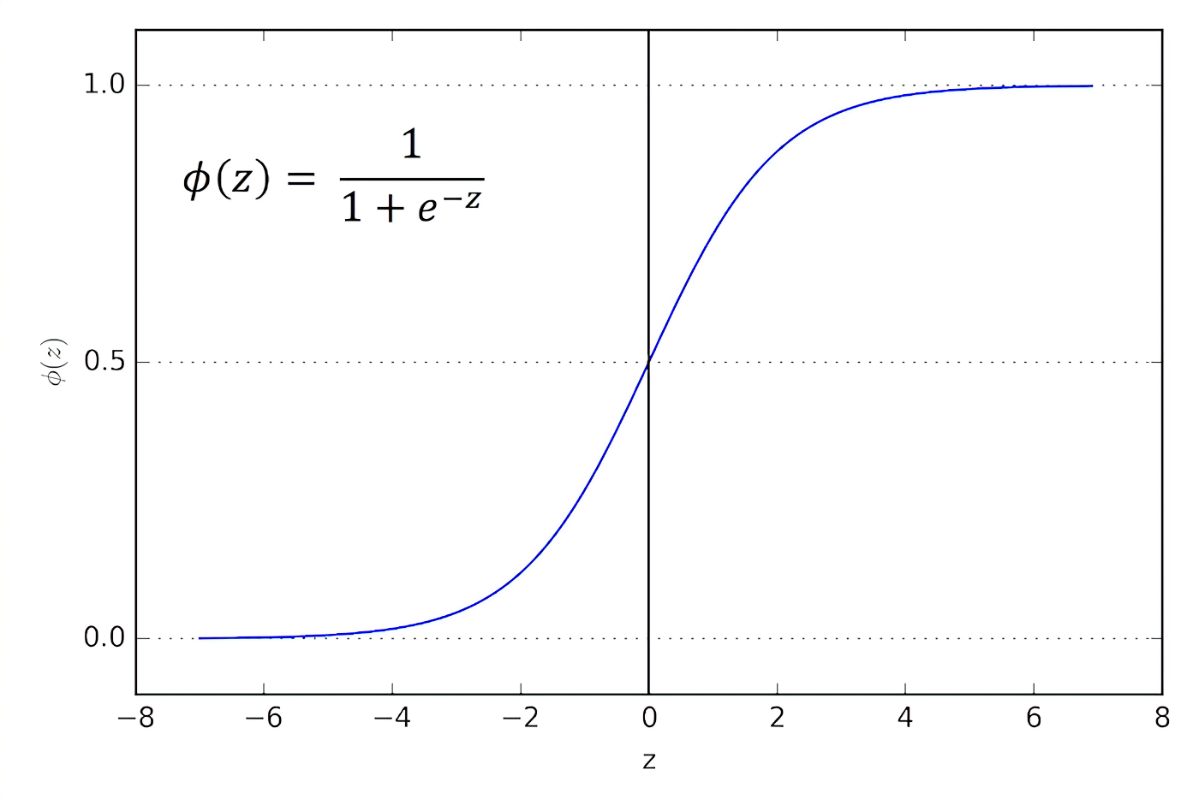
Analyze interactions
In xgboost, an interaction occurs when the downstream split has a higher gain than the upstream split.
# write the function collapse_tree to convert the tree output to interactions that occur in the tree.
collapse_tree <- function(t1){
t1 %>% group_by(Tree) %>% slice(which(Node == 0)) %>% ungroup %>%
select(Tree, Root_Feature = Feature) %>%
bind_cols(
t1 %>% group_by(Tree) %>% slice(which(Node == 1)) %>% ungroup %>%
select(Child1 = Feature)
) %>%
bind_cols(
t1 %>% group_by(Tree) %>% slice(which(Node == 2)) %>% ungroup %>%
select(Child2 = Feature)
) %>%
unite(col = "interaction1", Root_Feature, Child1, sep = ":", remove = F) %>%
select(-Child1) %>%
unite(col = "interaction2", Root_Feature, Child2, sep = ":", remove = T) %>%
pivot_longer(names_to = "names", cols = matches("interaction"), values_to = "interactions") %>%
select(-names)
}
xg_trees %>%
collapse_tree -> xg_trees_interactionsfind the top interactions in the model. The interactions are rated with different importance metrics, ordered by sumGain.
imps <- EIX::importance(xg1, hr1, option = "interactions")
as_tibble(imps) %>%
set_int(where(is.numeric))
#> # A tibble: 19 × 7
#> Feature sumGain sumCover meanGain meanCover frequency mean5Gain
#> <chr> <int> <int> <int> <int> <int> <int>
#> 1 satisfaction_level:n… 6818 24230 401 1425 17 691
#> 2 satisfaction_level:t… 5709 9017 713 1127 8 968
#> 3 average_montly_hours… 2794 2011 2794 2011 1 2794
#> 4 number_project:satis… 2688 3906 896 1302 3 896
#> 5 time_spend_company:s… 1991 2573 995 1286 2 995
#> 6 average_montly_hours… 1493 3264 298 652 5 298
#> 7 average_montly_hours… 1179 2483 392 827 3 392
#> 8 last_evaluation:time… 1171 2589 292 647 4 292
#> 9 last_evaluation:numb… 1003 3180 334 1060 3 334
#> 10 last_evaluation:sati… 904 6079 150 1013 6 163
#> 11 time_spend_company:l… 601 388 601 388 1 601
#> 12 last_evaluation:aver… 578 526 578 526 1 578
#> 13 number_project:last_… 431 1222 215 611 2 215
#> 14 satisfaction_level:a… 311 1772 155 885 2 155
#> 15 average_montly_hours… 290 606 290 606 1 290
#> 16 number_project:time_… 164 365 164 365 1 164
#> 17 Work_accident:time_s… 123 1144 123 1144 1 123
#> 18 satisfaction_level:l… 81 1115 81 1115 1 81
#> 19 time_spend_company:s… 75 1023 75 1023 1 75We can extract all the trees that contain the specified interaction.
imps[1,1] %>% unlist -> top_interaction
xg_trees_interactions %>%
filter(str_detect(interactions, top_interaction)) %>%
distinct -> top_interaction_trees
top_interaction_trees
#> # A tibble: 23 × 2
#> Tree interactions
#> <int> <chr>
#> 1 2 satisfaction_level:number_project
#> 2 4 satisfaction_level:number_project
#> 3 7 satisfaction_level:number_project
#> 4 11 satisfaction_level:number_project
#> 5 12 satisfaction_level:number_project
#> 6 16 satisfaction_level:number_project
#> 7 17 satisfaction_level:number_project
#> 8 18 satisfaction_level:number_project
#> 9 19 satisfaction_level:number_project
#> 10 20 satisfaction_level:number_project
#> # ℹ 13 more rowsThen extract the first 3 (most important) trees and print them.
top_interaction_trees$Tree %>% unique %>% head(3) -> trees_index
xgboost::xgb.plot.tree(model = xg1, trees = trees_index)We can confirm they are interactions because the child leaf in the interaction has higher split gain than the root leaf.
Analyze single features
# EIX package gives more detailed importances than the standard xgboost package
imps_single <- EIX::importance(xg1, hr1, option = "variables")
# choose the top feature
imps_single[1, 1] %>% unlist -> feature1
# get the top 3 rees of the most important feature. Less complicated than with interactions so
# no need to write a separate function like collapse tree
xg_trees %>%
group_by(Tree) %>%
slice(which(Node == 0)) %>%
ungroup %>%
filter(Feature %>% str_detect(feature1)) %>%
distinct(Tree) %>%
slice(1:3) %>%
unlist -> top_trees
xgboost::xgb.plot.tree(model = xg1, trees = top_trees)By looking at the 3 most important splits for satisfaction_level we can get a sense of how its splits affect the outcome.
shapley values
xg1 %>%
tidy_shap(hr1, form = hrf) -> hr_shaps
hr_shaps
#> $shap_tbl
#> # A tibble: 14,999 × 21
#> satisfaction_level last_evaluation number_project average_montly_hours
#> <dbl> <dbl> <dbl> <dbl>
#> 1 1.04 0.0302 1.30 0.0846
#> 2 -0.350 0.362 -0.294 0.213
#> 3 3.63 0.195 -0.264 0.142
#> 4 -0.380 0.364 -0.321 0.0841
#> 5 1.04 0.0302 1.30 0.0564
#> 6 1.04 0.0302 1.30 0.0846
#> 7 3.66 -0.147 -0.290 0.123
#> 8 -0.349 0.364 -0.294 0.213
#> 9 -0.326 0.797 -0.316 0.0841
#> 10 1.04 0.0302 1.30 0.0846
#> # ℹ 14,989 more rows
#> # ℹ 17 more variables: time_spend_company <dbl>, Work_accident <dbl>,
#> # promotion_last_5years <dbl>, sales_accounting <dbl>, sales_hr <dbl>,
#> # sales_it <dbl>, sales_management <dbl>, sales_marketing <dbl>,
#> # sales_product_mng <dbl>, sales_rand_d <dbl>, sales_sales <dbl>,
#> # sales_support <dbl>, sales_technical <dbl>, salary_high <dbl>,
#> # salary_low <dbl>, salary_medium <dbl>, `(Intercept)` <dbl>
#>
#> $shaps_long
#> # A tibble: 314,979 × 3
#> name SHAP FEATURE
#> <chr> <dbl> <dbl>
#> 1 (Intercept) -1.18 NA
#> 2 (Intercept) -1.18 NA
#> 3 (Intercept) -1.18 NA
#> 4 (Intercept) -1.18 NA
#> 5 (Intercept) -1.18 NA
#> 6 (Intercept) -1.18 NA
#> 7 (Intercept) -1.18 NA
#> 8 (Intercept) -1.18 NA
#> 9 (Intercept) -1.18 NA
#> 10 (Intercept) -1.18 NA
#> # ℹ 314,969 more rows
#>
#> $shap_summary
#> # A tibble: 21 × 5
#> name cor var sum sum_abs
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 (Intercept) NA 0 -17715. 17715.
#> 2 satisfaction_level -0.760 1.25 -2186. 12999.
#> 3 number_project -0.590 0.363 -2361. 8157.
#> 4 time_spend_company 0.711 0.408 -3111. 7693.
#> 5 last_evaluation 0.624 0.0555 -455. 3011.
#> 6 average_montly_hours 0.591 0.0329 -152. 1966.
#> 7 Work_accident -0.994 0.00728 -93.8 828.
#> 8 salary_low 0.958 0.00136 -28.7 531.
#> 9 promotion_last_5years NA 0 0 0
#> 10 salary_high NA 0 0 0
#> # ℹ 11 more rows
#>
#> $swarmplot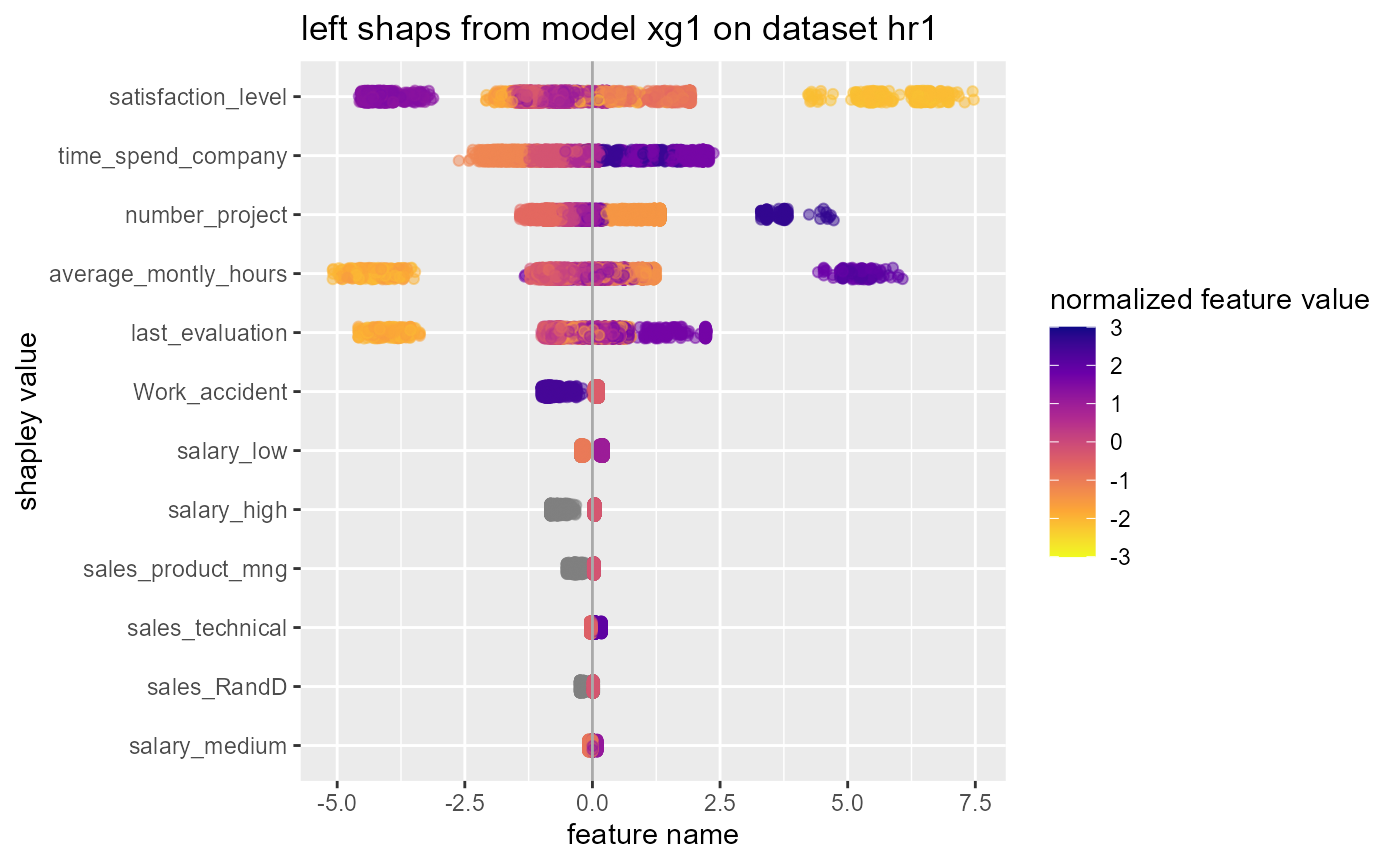
#>
#> $scatterplots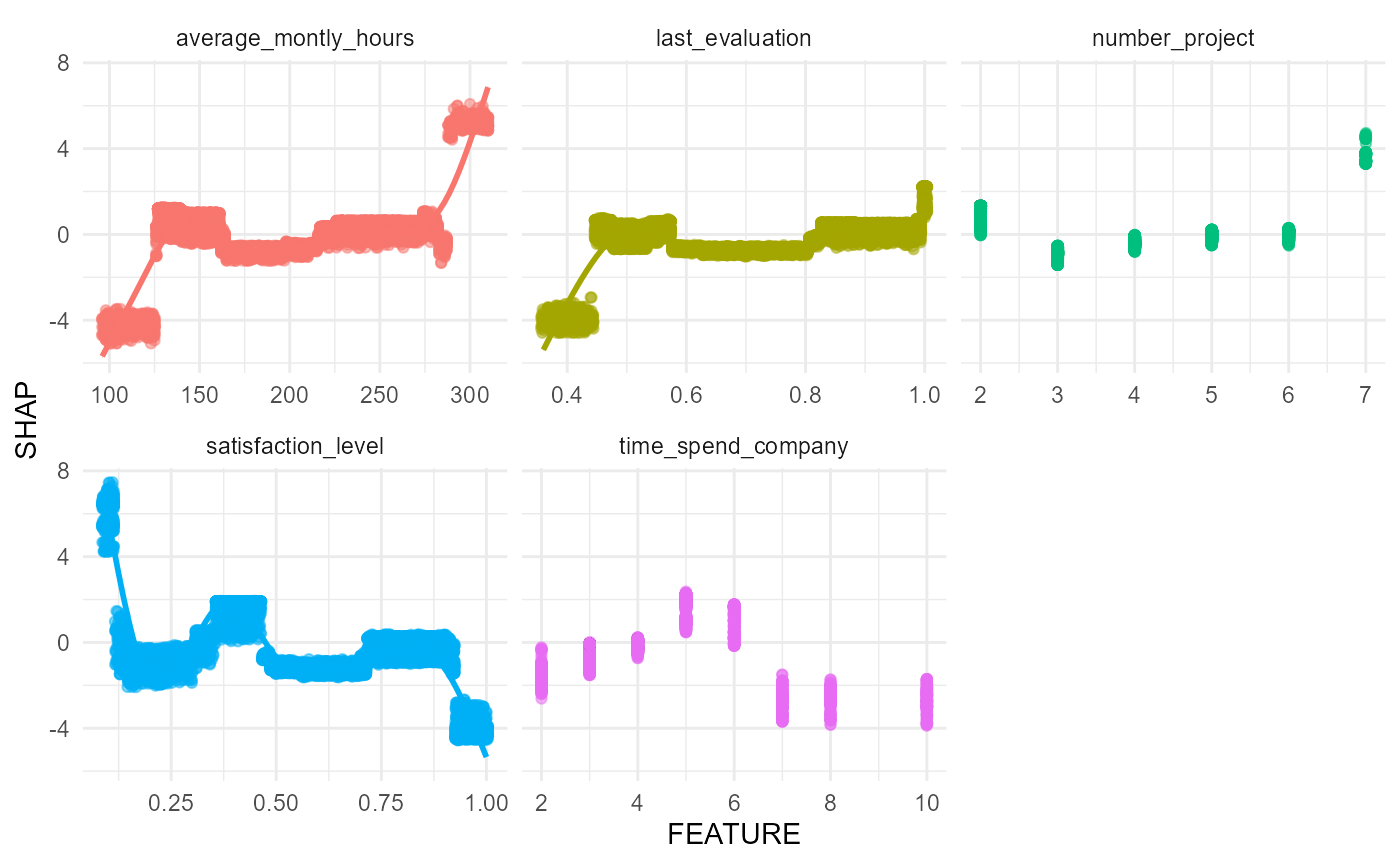
#>
#> $boxplots